 Life History Traits and Vegetation Memory Components
Life History Traits and Vegetation Memory Components
Disclaimer
This project is currently on hold.
Motivation
Throughout my previous project concerning Vegetation Memory across Global Dryland Regions, I identified prominent spatial patterns of intrinsic and extrinsic vegetation memory components. These patterns may serve us in discerning global and local resistance and recovery potential of vegetation communities to perturbations.
However, these patterns alone only tell part of the story. How do these come about? How are they maintained? Understanding these mechanisms would undoubtedly go a long way in predicting responses of vegetation to anthropogenic and climate change-driven disturbances. Ultimately, I expect this exercise to create valuable knowledge for conservation biology and the agricultural sector.
Description
MODIS EVI data is aggregated at bi-weekly intervals and used as a proxy of vegetation response in the dryland study regions between January 2000 and December 2019. Independent climate data is provided using the KrigR package (
KrigR - Downloading and Downscaling of ERA5(-Land) data using R). Doing so allows us to make use of the high temporal resolution of the European Centre for Medium-range Weather Forecasts ReAnalysis 5 (ERA5) data from the European Centre for Medium-Range Weather Forecasts (ECMWF) at spatial resolutions of roughly 1x1km. Effectively, this increases the spatial resolution of vegetation memory products by almost one order of magnitude when compared to my previous project at 9x9km resolution. Vegetation memory is calculated the same way as during my previous project.
Using these high-resolution vegetation memory products, we can reasonably argue that ground-data obtained through functional trait campaigns or vegetation plot exercises represents the 1x1km grid reasonably well. Thus we are able to assess correlations between the different vegetation components and ground-truthed expressions of plant life. To identify what shapes vegetation memory expressions, I am investigating a range of three potential predictor families:
- Plant Functional Traits - I expect investment in certain functional characteristics to greatly influence resistance and recovery potential. For example, sturdy leaves should grant resistance to temperature fluctuations but are costly to recover once lost.
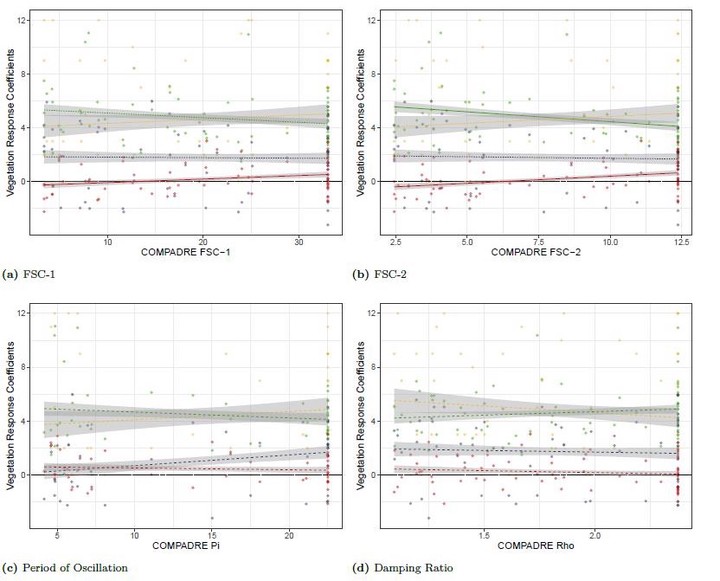
- Life History Traits - Life history speed and timing undoubtedly alters vegetation memory expressions. But which life history traits fair best at explaining these processes? For example, a short-lived population may be less resistant to a perturbation than a long-lived one, but surely boasts a reproduction rate that achieves fast recovery.
- Abiotic Legacies - Biological organisms adapt to their surroundings. On varying time-scales and through varying processes, but they adapt nonetheless. I aim to investigate how strong these adaptations become by computing the impact of the legacy mean and standard deviation of abiotic conditions on vegetation memory components. For example, a species in a highly variable aridity regime may be more resistant to fluctuations in aridity than one used to stable, moist conditions